library(MASS)A Note on Projection Matrices (and an AI Attempt to Create Visualizations)
I am sharing my 50-minute use of AI to create a set of lecture notes to help students understand projection matrices. The visualization part was where I had hoped AI would save time. However, that turned out to be the most problematic part - AI either did not fully understand the task or did not know how to implement it.
I have omitted a lengthy and unsuccessful exchange. In the end, I tried Gemini; perhaps because I finally provided enough details, I was able to obtain the plot I wanted.
I have created similar visualizations manually before. If I had revisited my old code and adapted it for this note, it likely would have taken a similar amount of time—or even less.
Outline (Written by Yu)
- What is the definition of a projection matrix?
- What are the properties of a projection matrix?
- How does a projection matrix look like?
- What does a projection matrix do? In a 2D dimension, what does a projection matrix do to data?
- Simulate 20 data points from a bivariate normal with mean (1,1), variances 1 and covariance 0.7. We examine five projections:
- to the x-axis,
- to the y-axis
- \(y=x\),
- \(y=2x\).
- Simulate 20 data points from a bivariate normal with mean (1,1), variances 1 and covariance 0.7. We examine five projections:
Definition (Written by Copilot)
- A projection matrix \(\mathbf P\) is a square matrix that satisfies the following condition: \[\mathbf P^2=\mathbf P\]
- The projection matrix \(\mathbf P\) is said to project onto the column space of \(\mathbf P\).
Yu: It is incorrect. The definition of a projection matrix is not just that it is idempotent. It also needs to be symmetric.
Properties(Written by Copilot)
- A projection matrix \(\mathbf P\) is idempotent, i.e., \(\mathbf P^2=\mathbf P\).
- A projection matrix \(\mathbf P\) is symmetric, i.e., \(\mathbf P^T=\mathbf P\).
- The eigenvalues of a projection matrix \(\mathbf P\) are either 0 or 1.
- The rank of a projection matrix \(\mathbf P\) is equal to the trace of \(\mathbf P\), i.e., \(\text{rank}(\mathbf P) = \text{tr}(\mathbf P)\).
- The projection matrix \(\mathbf P\) can be expressed as \(\mathbf P = \mathbf A(\mathbf A^T\mathbf A)^{-1}\mathbf A^T\), where \(\mathbf A\) is a matrix whose columns form a basis for the column space of \(\mathbf P\).
Examples (Written by Copilot)
- The projection matrix that projects onto the x-axis is given by \(\mathbf P = \begin{bmatrix} 1 & 0 \\ 0 & 0 \end{bmatrix}\).
- The projection matrix that projects onto the y-axis is given by \(\mathbf P = \begin{bmatrix} 0 & 0 \\ 0 & 1 \end{bmatrix}\).
- The projection matrix that projects onto the line \(y=x\) is given by \(\mathbf P = \frac{1}{2}\begin{bmatrix} 1 & 1 \\ 1 & 1 \end{bmatrix}\).
- The projection matrix that projects onto the line \(y=2x\) is given by \(\mathbf P = \frac{1}{5}\begin{bmatrix} 1 & 2 \\ 2 & 4 \end{bmatrix}\).
Simulation (Written by Copilot)
Code
set.seed(123)
n <- 20
mu <- c(1, 1)
sigma <- matrix(c(1, 0.7, 0.7, 1), nrow = 2)
data <- mvrnorm(n, mu, sigma)
# Define projection matrices
P_x <- matrix(c(1, 0, 0, 0), nrow = 2)
P_y <- matrix(c(0, 0, 0, 1), nrow = 2)
P_xy <- matrix(c(0.5, 0.5, 0.5, 0.5), nrow = 2)
P_2xy <- matrix(c(1/5, 2/5, 2/5, 4/5), nrow = 2)Apply projection matrices to data
Code
data_x <- data %*% P_x
data_y <- data %*% P_y
data_xy <- data %*% P_xy
data_2xy <- data %*% P_2xyPlot original and projected data
Code
par(mfrow=c(2,3))
plot(data, main="Original Data", xlim=c(-1,3), ylim=c(-1,3))
plot(data_x, main="Projection to x-axis", xlim=c(-1,3), ylim=c(-1,3))
plot(data_y, main="Projection to y-axis", xlim=c(-1,3), ylim=c(-1,3))
plot(data_xy, main="Projection to line y=x", xlim=c(-1,3), ylim=c(-1,3))
plot(data_2xy, main="Projection to line y=2x", xlim=c(-1,3), ylim=c(-1,3))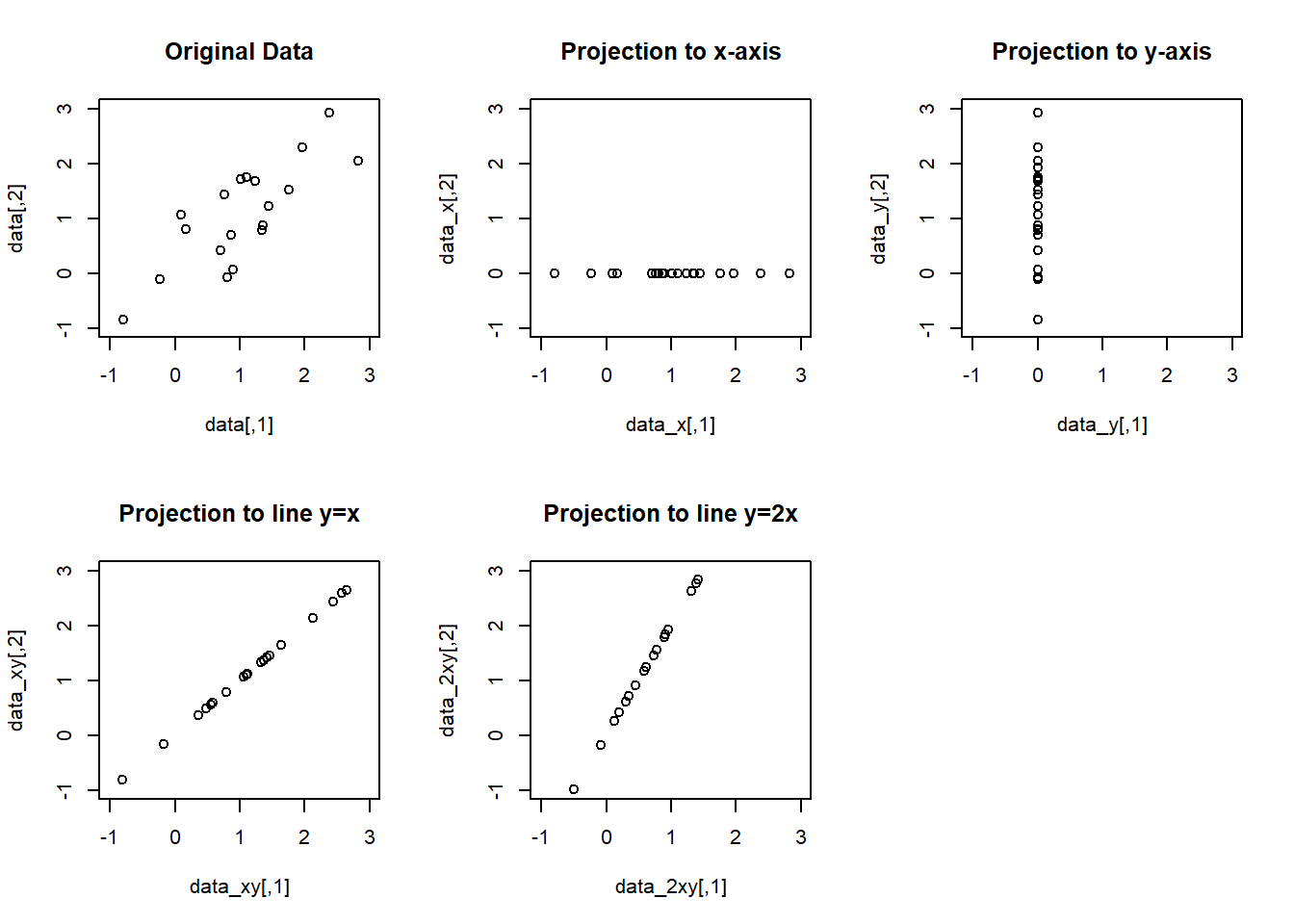
Redraw the data
Yu: Redraw the data so that, for each projection, the projected data are shown using little ticks and the original data are shown in points. They need to be in the same plot.
Copilot:
Code
par(mfrow=c(1,1))
plot(data, main="Projection to x-axis", xlim=c(-1,3), ylim=c(-1,3))
points(data_x, col="red", pch=16)
points(data_y, col="blue", pch=16)
points(data_xy, col="green", pch=16)
points(data_2xy, col="purple", pch=16)
legend("topright", legend=c("Original Data", "Projection to x-axis", "Projection to y-axis", "Projection to line y=x", "Projection to line y=2x"),
col=c("black", "red", "blue", "green", "purple"), pch=16)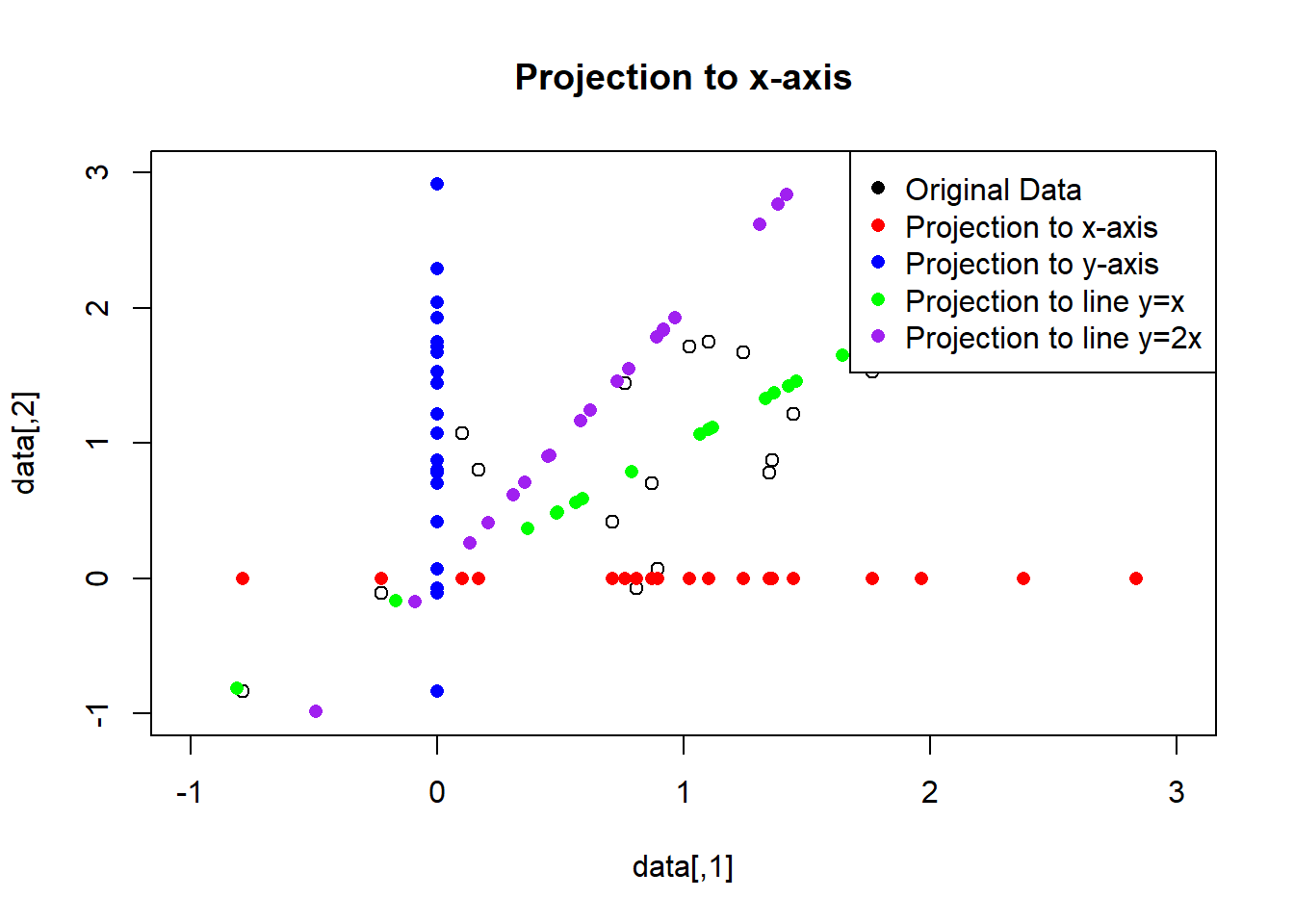
Redraw the data with ticks
Yu: Still not what I want. The ticks are short bars of the same length. Each bar should be perpendicular to the space projected to. For example, the data projected to the x-axis should be shown using vertical ticks, i.e., ticks perpendicular to the x-axis.
Gemini:
Code
plot(data, main="Projection to x-axis", xlim=c(-1,3), ylim=c(-1,3))
# Function to add ticks for projected data
add_ticks <- function(data, projection, col) {
for (i in 1:nrow(data)) {
x0 <- data[i, 1]
y0 <- data[i, 2]
x1 <- projection[i, 1]
y1 <- projection[i, 2]
segments(x0, y0, x1, y1, col=col)
}
}
add_ticks(data, data_x, col="red")
add_ticks(data, data_y, col="blue")
add_ticks(data, data_xy, col="green")
add_ticks(data, data_2xy, col="purple")
legend("topright", legend=c("Original Data", "Projection to x-axis", "Projection to y-axis", "Projection to line y=x", "Projection to line y=2x"),
col=c("black", "red", "blue", "green", "purple"), pch=16)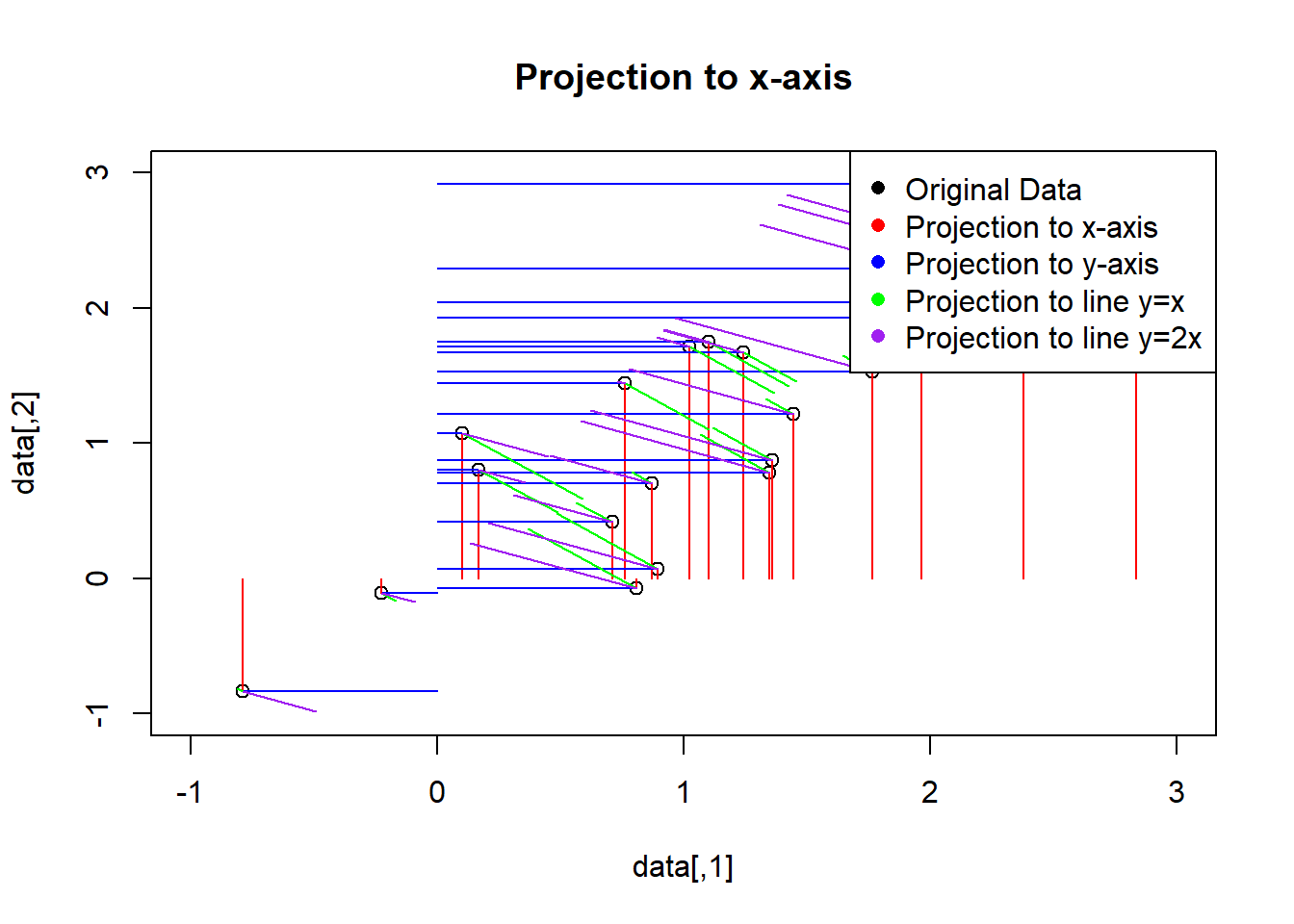
Still incorrect. The bars are of varying lengths.
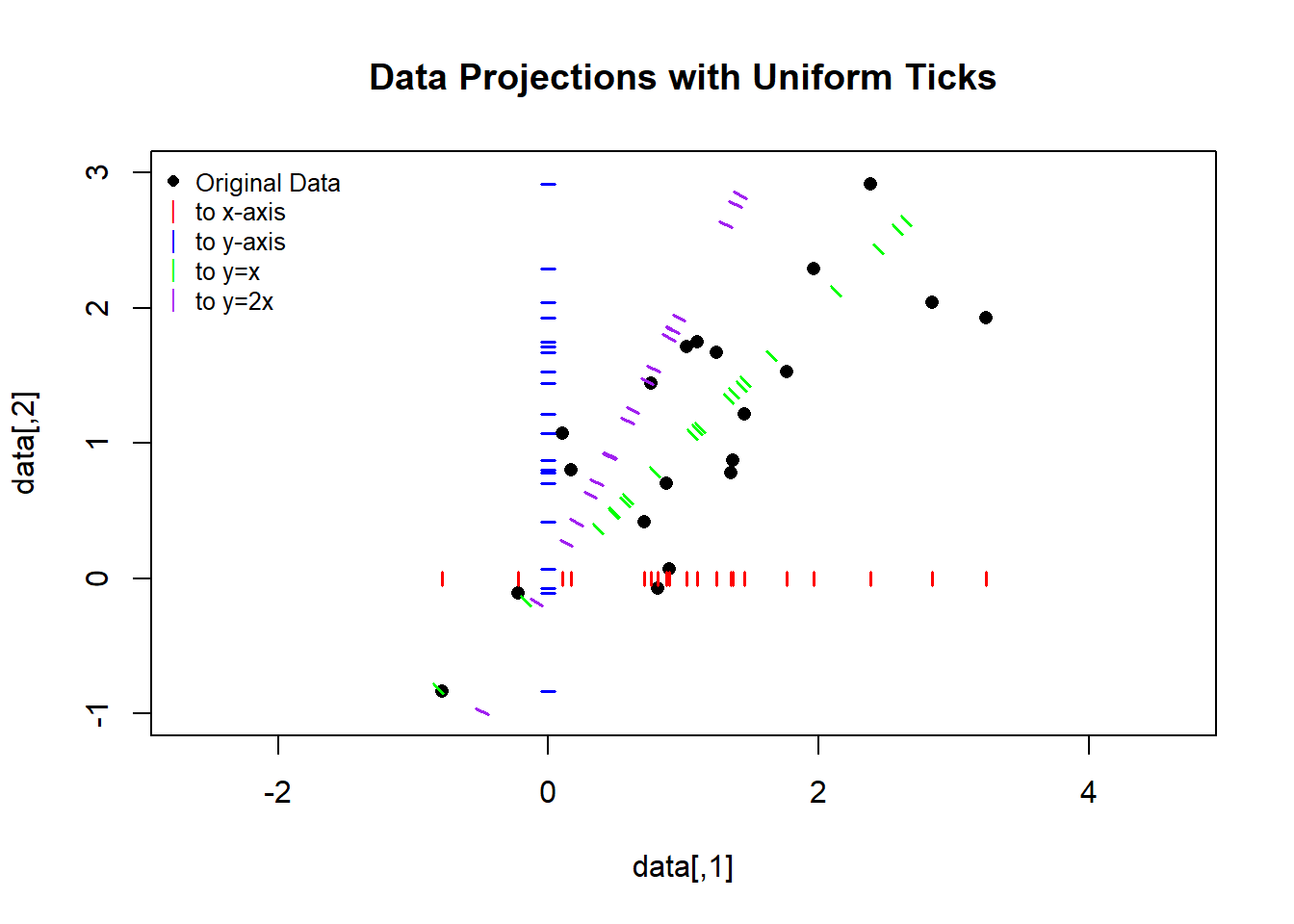
plot(data, main = "Data Projections with Uniform Ticks",
xlim = c(-1, 3), ylim = c(-1, 3), pch = 16, asp = 1)
# Function to add uniform, perpendicular ticks
# target_vec is a vector defining the line projected onto (e.g., c(1,0) for x-axis)
add_ticks <- function(projected_data, target_vec, col, tick_len = 0.1) {
# Get the perpendicular vector (rotate target_vec by 90 degrees)
# For a vector (a, b), the perpendicular is (-b, a)
perp_vec <- c(-target_vec[2], target_vec[1])
# Normalize the perpendicular vector to unit length
perp_vec <- perp_vec / sqrt(sum(perp_vec^2))
for (i in 1:nrow(projected_data)) {
px <- projected_data[i, 1]
py <- projected_data[i, 2]
# Calculate start and end points of the tick centered at the projected point
x0 <- px - (perp_vec[1] * tick_len / 2)
y0 <- py - (perp_vec[2] * tick_len / 2)
x1 <- px + (perp_vec[1] * tick_len / 2)
y1 <- py + (perp_vec[2] * tick_len / 2)
segments(x0, y0, x1, y1, col = col, lwd = 1.5)
}
}
# 1. Projection to x-axis: line is (1,0)
add_ticks(data_x, c(1, 0), col = "red")
# 2. Projection to y-axis: line is (0,1)
add_ticks(data_y, c(0, 1), col = "blue")
# 3. Projection to y=x: line is (1,1)
add_ticks(data_xy, c(1, 1), col = "green")
# 4. Projection to y=2x: line is (1,2)
add_ticks(data_2xy, c(1, 2), col = "purple")
legend("topleft",
legend = c("Original Data", "to x-axis", "to y-axis", "to y=x", "to y=2x"),
col = c("black", "red", "blue", "green", "purple"),
pch = c(16, 124, 124, 124, 124), bty = "n", cex = 0.8)